Analyzing Antibiotic Resistance, Lab Report Example
Analyzing Antibiotic Resistance to Gram-Positive and Gram-Negative Bacteria
Abstract
There are many different types of bacteria and not every antibiotic is effective against every bacteria. Since there is such a variation of different types of bacteria, antibiotics are usually made specifically for a specific type of bacteria. The purpose of this laboratory exercise was to demonstrate the effect Penicillin, Ampicillin, Erythromycin, and Tetracylcine on the Gram-positive bacteria Escherichia Coli and Pseudomonas Aeruginosa and the Gram-negative bacteria Bacillus Cereus and Staphylococcus Aureus. These results indicated that the Staphylococcus Aureus was susceptible to each antibiotic tested and Pseudomonas Aeruginosa was resistant to each antibiotic tested. Pseudomonas Aeruginosa was the only bacteria that were resistant to all the types of antibiotics used, which correlates previous research indicate its resistance to antibiotics through plasmids. E. coli was also resistant to Penicllin and have plasmids that carry genetic information for antibiotic resistance, like P. aeuruginosa. Results of this study were accurate per the standardized methods
Introduction
Antibiotics are a type of antimicrobial agent developed from microorganisms such as bacteria and fungi. Antibiotics were developed in order to fight bacteria and are used in clinical medicine as well as in agriculture. There are many different types of bacteria and not every antibiotic is effective against every bacteria. Antibiotics are characterized according to their molecular make-up, type of microbial action, and spectrum of antimicrobial behavior. In fact, according to Madigan et al., (2012) there are more than 10,000 metric tons of different types of antimicrobial drugs developed and used each year due to the different species of bacteria. Since there is such a variation of different types of bacteria, antibiotics are usually made specifically for a specific type of bacteria. There are, however, antibiotics known as “wide spectrum” antibiotics that can be used to fight more than one type of bacteria. Table 1 illustrates different types of antibiotics and the types of bacteria in which cell growth is prevented. As seen, the antibiotic Isoniazid only helps prevent the growth of one genus of bacteria, the Mycobacteria; whereas Tetracycline helps prevent the growth of gram-negative bacteria, gram-positive bacteria, and parasitic bacteria. The purpose of this laboratory exercise was to demonstrate the effect of four different types of antibiotics on two different types of bacteria: Gram-positive and Gram-negative. The Gram-positive bacteria consisted of Escherichia Coli and Pseudomonas Aeruginosa and the Gram-negative bacteria consisted of Bacillus Cereus and Staphylococcus Aureus. The types of antibiotics used were Penicillin, Ampicillin, Erythromycin, and Tetracylcine.
Gram-negative and Gram-positive bacteria were chosen for this exercise because, in microbiology, the Gram-stain is usually the first step in classifying and characterizing bacteria. (Madian et al., 2012) These particular bacteria also have known differences in susceptibilities to different antibiotics. In addition, gram-positive and gram-negative bacteria differ in their response to different types of antibiotics, such as penicillin. Gram-positive bacteria, for instance, are usually affected by the penicillin while gram-negative bacteria are not affected. Penicillin is an antibiotic that was first developed in 1929 by Alexander Fleming. It was isolated from Penicillium chrysogenum, a fungus, and was the first clinically useful antibiotic on the market. (Madigan et al., 2012) Penicillin works in inhibiting cell wall synthesis of bacteria, thereby deterring their growth.
Erythromycin is an antibiotic that is used in individuals who are allergic to penicillin. This antibiotic is a wide-spectrum antibiotic that is produced from a strain of Streptomyces, a type of bacteria. Erythromycin is active against both gram-positive and gram-negative bacteria and is usually used to treat against Mycoplasmas, a genus of bacteria that lack a cell wall, respiratory tract infections, Chlamydia, Syphilis and Legionnairs’ disease. (Kanfer et al.) Ampicillin is a derivative of penicillin that works by inhibiting the peptidoglycan chains in the cell wall of eubacteria, causing cells growing in ampicillin to burst. (Russell and Chopra, 1990) Ampicillin is effective against some gram-positive and gram-negative bacteria such as Streptococcus, Enterococcus, Listeria, influenza, Salmonella and Shigella. (Antimicrobe, 2012) Also effective against both Gram positive and Gram negative bacteria is the the antibiotic Tetracycline.. Tetracycline was also developed from Streptomyces and was the first wide-spectrum antibiotic. It works in inhibiting protein synthesis of bacteria.
In addition to the antibiotics used in this lab, there were some common types of infectious bacteria used as well. Escherichia coli causes infectious diseases that are obtained from fecal contaminated foods. It is the most widely studied prokaryotic organism because it can be easily and inexpensively grown in a laboratory setting. Staphylococcus aureus can cause a range of illnesses, from minor skin infections such as pimples and impetigo, to life-threatening diseases such as pneumonia, meningitis and toxic shock syndrome. Bacillus cereus is responsible for some foodborne illnesses which cause nausea, vomiting and diarrhea. Pseudomonias aeruginosa typically infects the pulmonary tract, urinary tract, burns, wounds, and also causes other blood infections.
Antibiotics that were at one time effective in controlling many of these infections are not effective anymore due to the resistance acquired from the bacteria. This type of resistance is found in both veterinary and human medicine. (Salehi and Bonab, 2006) These antibiotics have been used so widely and for so long that the infectious organisms they are designed to kill have adapted to them, making them less effective. To deal with this acquired resistance, new antimicrobial compounds are being developed in several ways, including new analogs of existing antimicrobial compounds and computer aided drug design.
In this lab, the purpose is to see which antibiotics inhibit growth of bacteria. Antibiotics such as penicillin should work better on the gram-positive bacteria due to their thicker cell wall. Ampicillin, a semi synthetic penicillin should be able to work on either the gram-positive or gram negative bacteria because its development offers a broader range of activity. Some of the wide-spectrum antibiotics that inhibit protein synthesis, such as erythromycin, should work better on the gram-positive bacteria. Erythromycin is often used in patients who are allergic to penicillin. (Madigan et al., 2012) The antibiotic produced from the Streptomycin, tetracycline, is one of the first broad-spectrum antibiotics, inhibiting most gram positive and gram negative bacteria. It is useful in treating infections of the urinary tract, chlamydia, Rickettsia and Lyme disease.
In order to compare the effects of bacteria on different antibiotics and measure the degree of resistance, the Kirby-Bauer test was conducted and the results were analyzed and compared to a standardized table. It is therefore predicted that if a bacterial colony is susceptible to an antibiotic, then a zone of inhibition will be created around the bacteria disc, indicating no growth. If bacteria are resistant to the antibiotic, then there will be no zone of inhibition present around the bacteria disc. If a bacteria is slightly susceptible to the antibiotic, then the zone of inhibition will measure slightly in-between the bacteria that were susceptible and resistant.
Materials and Methods
Agar plates were obtained and carefully labeled per type of bacteria. In addition, the students initials were placed on the edge of the plate. A sterile dry cotton swab was dipped into the bacteria broth and the agar plate was inoculated in three directions in order to spread the bacteria evenly over the agar surface. The cotton swab was discarded into an appropriate container. Filter paper antibiotic discs were placed on the surface of the agar using tweezers disinfected with alcohol. The petri dishes were then left in the laboratory to allow for bacterial growth.
During the following laboratory, the petri dishes were examined. Petri dishes that had clear areas, where bacteria did not grow around the antibiotic discs, were referred to as zones of inhibition. The diameter of each zone of inhibition was measured in millimeters using a ruler and the results were recorded. The zone diameter was compared to a standardized chart from Madian et al., (2012) textbook and analyzed for susceptibility. Results were then put into table format.
Results
The results of the study were documented and compared using a standardized chart (Table 2 and Table 3). These results indicated that the Staphylococcus Aureus was susceptible to each antibiotic tested. The Pseudomonas Aeruginosa was resistant to each antibiotic tested. (Table 2) Escherichia Coli was resistant to Penicillin only, susceptible to Ampicillin and Tetracycline and intermediate to Erythromycin. Bacillus Cereus was resistant to both Penicillin and Ampicillin and susceptible to both Erythromycin and Tetracycline. In addition, the bacteria with the largest diameter of zone of inhibition were Staphylococcus Auerus tested with Penicillin. The bacteria with the smallest recorded diameter of zone of inhibition were Pseudomonas Aeruginosa with Penicillin, Ampicillin and Erythromycin, all at 6.0 mm and all resistant.
Table 1. Represents the antimicrobial spectrum of action. Different antibiotics affect a specific group of bacteria. A few antibiotics area very specific and only affect one species of bacteria. (Madigan et al., 2012)
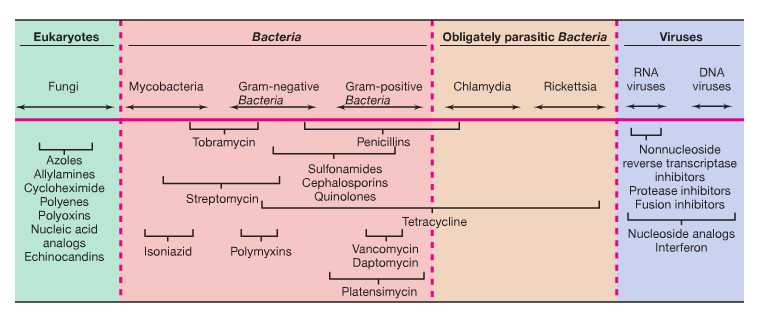
Table 2. Antibiotic susceptibility results from two Gram Positive Bacteria (E. Coli and P. Aeruginosa) and two Gram Negative Bacteria (B. Cereus and S. Aureus) tested against four different types of Antibiotics.
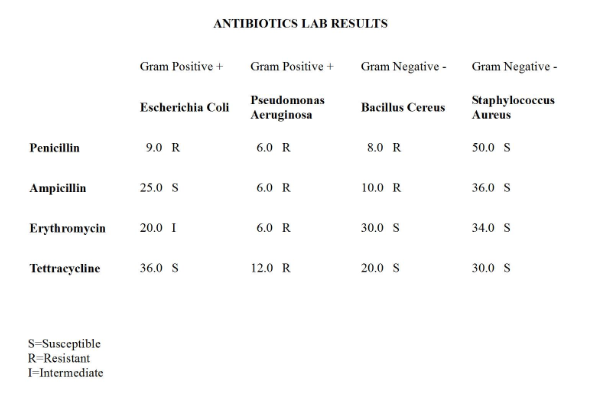
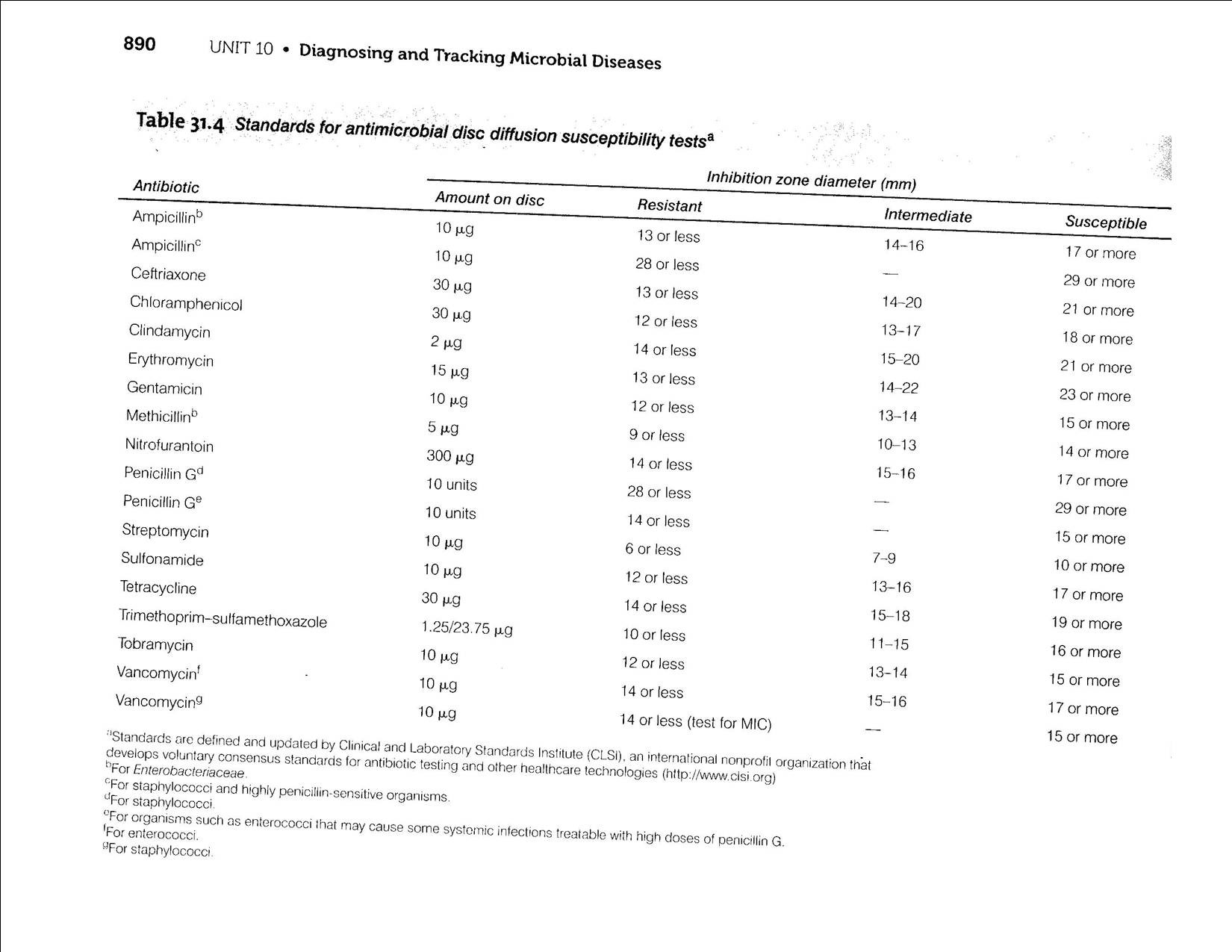
Table 3. Standard table for antimicrobial disc diffusion susceptibility test (Madian et al., 2012)
Discussion
Different types of antibiotics inhibit different types of bacterial growth depending on the action of the antiobiotic. For instance, Penicillin can only prevent gram-positive bacteria from growing because it prevents cell wall growth; whereas, other antibiotics interfere with nucleic acid synthesis or metabolite synthesis and can therefore be used on various types of bacteria. (Westminster College) Widespread antibiotic use is one of the most important factors in the occurrence of antibiotic resistance in microorganisms. Treatment with antibiotics usually occurs through the use of daily doses for a specific amount of time. The treatment of antibiotics eradicates the bacteria and prevents further growth or emergence of the bacteria. However, failure to follow the proper treatment plan of antibiotics can cause the bacteria to reactivate and become resistant to the original antibiotic. This resistance must be tested in order to determine the antibiotic susceptibility. In this study, four different bacteria were tested against four different antibiotics. The results were compared to a standarized chart and termed either suscpetible, intermediate or resistant. One bacteria, Staphylococcus Aureus, was susceptible to each antibiotic tested. The results of this study are accurate with previous research. Staphylococcus aureus is known to be naturally susceptible to every antibiotic that has been produced due to the transfer of genes. However, some strains of S. Aureus have been becoming resistant to antibiotics (Chambers and DeLeo, 2009) . Nevertheless, the susceptibility of S. aureus is what led Fleming to the discovery of penicillin. (Chambers and DeLeo, 2009) Penicillin was the first antibiotic; however, after a few years of its discovery, penicillin began to become resistant. In this study, three types of bacteria were resistant to Penicillin, both Gram-positive and one Gram-negative bacteria. The interesting result in this study was that the Gram-positive bacteria were resistant to the penicillin. In general, Gram-positive bacteria are usually susceptible; however, there are some Gram-positive bacteria that are resistant to penicillin, such as Bacillus and Staphylococcus. In a study conducted by Andrews and Wise (2002), all five isolates of Bacillus cereus tested were resistant to pencillin in the presence of ß-lactamase.
Pseudomonas Aeruginosa, Gram-negative bacteria, was the only bacteria that were resistant to all the types of antibiotics used. The results of this study correlate with known research, that the P. aeurginosa is naturally resistant to antibiotics. Infections caused by these bacteria are difficult to treat because resistance is caused from the resistance transfer plasmid or R plasmid, which carry genes that remove antibiotics from their cell. (Madian et al., 2012 ) Basically, P. aeurginosa develops resistance to antibiotics through its ability to acquire resistance genes on genetic structures known as plasmids, or through mutations that change gene expression or function. Due to their ability to undergo these strategies they have become resistant and limit treatment options with antibiotics. (Lister et. Al, 2009)
Escherichia Coli was resistant to Penicillin only, susceptible to Ampicillin and Tetracycline and Intermediate to Erythromycin. E. coli contain plasmids that carry genetic information for antibiotic resistance, like P. aeuruginosa they are able to acquire the ability to become resistant. Findings from a study conducted by Schroeder et al. (2002) indicated that antibiotic resistance was widespread in E. coli that inhabit humans and food animals and linked these findings to the plasmids in E. coli.
Since Penicillin has been used for many years, it has allowed for E. coli to develop genes for resistance. The antibiotic resistance genes are located on plasmids and can move from cell to cell as well as from one bacteria species to another bacteria species. Therefore, it is common for these bacteria to be resistant to commonly used antibiotics. Another example of this resistance was observed in P. aerginosa and its resistance to tetracycline. Although tetracycline is supposed to have effects on both Gram-positive and Gram-negative bacteria, it has been used extensively in the medical field and has therefore bacteria have become extremely resistant to this antibiotic. (Madian et al., 2012)
There was a difference in the diameter of zone of inhibition between the Gram-Positive bacteria and Gram-negative bacteria. The largest zone of inhibition was observed in the Gram-negative bacteria and the smallest zone of inhibition was observed in the Gram-Positive bacteria, most likely due to their thicker cell wall. Penicillin and ampicillin both targets the inhibition of cell wall synthesis, with Penicillin, as mentioned previously becoming a bacteria-resistant antibiotic. The results confirm this, showing that the largest zone of inhibition is seen in the gram-negative bacteria, which, as previously mentioned, are comprised of a different type of cell wall than gram-positive bacteria.
Many of the bacteria acquire resistance to antibiotics through destruction of the drug, prevention of the drug target site, mutation of the drug target site and genetic mutations. (Kardar) In addition, there are human methods that have increased the resistance to antibiotics, such as not completing the required length or dose of antibiotics. The results of this study show that there are bacteria, both Gram-positive and Gram-negative that are still susceptible to antibiotics, but are also resistant as well. There are some errors or differences that could have occurred during this study and altered results, such as the concentration of the antibiotics on these disks, temperature of the incubation, as well as the time of incubation and thickness of the agar plates; however, the bacteria used were standardized and the results of this study were accurate.
References
Andrews, J.M. and Wise, R. Susceptibility testing of Bacillus species. (2002). J. Antimicrob Chemother. 49:1040:1042.
Antimicrobe. (2012). Infectious Disease Microbial Agents. Retrieved from: http://www.antimicrobe.org/drugpopup/ampicillin.htm
Chambers, H. and DeLeo, F.R. (2009). Waves of Resistance: Staphylococcus aureus in the Antibiotic Era. Nat Rev Microbiol. 7(9):629-641.
Kanfer, I., Skinner M.F., Walker, R.B. ( ). Analysis of macrolide antibiotics.
Kardar, S.S. Antibiotic Resistnace: New Approaches to a Historical Problem. An ActionBioscience.org original article: http://www.actionbioscience.org/newfrontiers/kardar.html
Lister, P.D., Wolter, D.J, Hanson, N.D. Antibacterial-Resistant Pseudomonas aeruginosa: Clinical Impact and Complex Regulation of Chromosomally Encoded Resistance Mechanisms. (2009) Clinical Microbiology. 22(4):582-610.
Madigan, M.T., Martinko, J.M., Stahl, D.A., Clark, D.P. (2012). Madian et al., 2012 Biologyof Microorganisms. Benjamin Cummings Publishing, San Francisco CA. 1004pp.
Russell, A., and I. Chopra. 1990. Understanding Antibacterial Action and Resistance. Ellis Horwood, NY.
Salehi, T.Z. and Bonab, S.F. (2006). Antibiotics Susceptibility Pattern of Escherichia coli Strains Isolated from Chickens with Colisepticemia in Tabriz Province, Iran. International Journal of Poultry Science 5 (7): 677-684, 2006.
Schroeder CM, Meng J, Zhao S, DebRoy C, Torcolini J, Zhao C, et al. Antimicrobial resistance of Escherchia coliO26, O103, O111, O128, and O145 from animals and humans. Emerg Infect Dis [serial online] 2002 Dec [date cited]. http://dx.doi.org/10.3201/eid0812.020070
Westminster College. Antibiotic Inhibition of Bacteria. Retrieved from: http://www.westminster.edu/acad/sim/pdf/santibioticinhibitionofbacteria.pdf.

Time is precious
don’t waste it!

Plagiarism-free
guarantee

Privacy
guarantee

Secure
checkout

Money back
guarantee






